View sample Y-Chromosomes and Evolution Research Paper. Browse other research paper examples and check the list of research paper topics for more inspiration. If you need a religion research paper written according to all the academic standards, you can always turn to our experienced writers for help. This is how your paper can get an A! Feel free to contact our custom writing service for professional assistance. We offer high-quality assignments for reasonable rates.
For several decades now, genetic studies have been used as a complement to more traditional anthropological approaches in the study of human origins and diversification. Most genetic analyses have used autosomal and mitochondrial (mtDNA) markers and such data provide a global picture of human evolution (Cavalli-Sforza et al. 1994, Ruiz-Linares 1999). Two major findings are that African populations show a higher level of genetic diversity when compared with other continents and that the greatest differentiation between human populations is seen between African and non-African populations. Both these observations are in agreement with paleoanthropological data showing that humans originated in Africa. Interestingly, the overall level of genetic differentiation between human populations is small relative to the diversity observed within populations, suggesting that the differentiation of human populations occurred relatively recently, possibly around 100,000–200,000 years ago. Based on paleoanthropological evidence two general models of human origins have been proposed: a multiregional and a Garden of Eden model. Under the first scenario the diversification of human populations is very old, tracing back to the initial differentiation of regional Homo erectus populations. The Garden of Eden model proposes that the differentiation of human population is recent (of the order of 100,000 years) and dates to the migration out of Africa of a fully modern human species. Genetic data have been found to be mostly consistent with a recent out-of-Africa model.
Academic Writing, Editing, Proofreading, And Problem Solving Services
Get 10% OFF with 24START discount code
Until recently, little was known about the extent and distribution of diversity in the Y-chromosome. This problem stemmed from a basic difficulty in identifying genetic variation on this chromosome. None of the traditional genetic markers are found on the Y and attempts at identifying molecular variation progressed slowly until the late 1990s. A major breakthrough came about in 1997 when a novel technique known as Denaturing High Pressure Liquid Chromatography (DHPLC) was developed and allowed the rapid screening of a large number of DNA fragments from the Y-chromosome, leading to an increase in the speed of identification of new molecular variants. Recent developments resulting from the Human Genome Project, such as the availability of a draft sequence of the human genome and the activities of the so-called SNP (single nucleotide polymorphism) consortium are leading to an even faster identification of molecular variants on the Y-chromosome. The recent availability of a large number of genetic markers on the Y-chromosome has led to an increasing interest in the use of this chromosome in human evolutionary studies, both to explore human origins and ancient diversification as well as to examine recent population history.
1. Transmission Of The Y-Chromosome
Part of what makes the Y-chromosome of special interest for human population genetic studies is its peculiar mode of transmission. The passage of the Y from one generation to the next is immediately apparent: it determines the male sex. In other words, it is only transmitted from fathers to their male offspring. The fact that the Y-chromosome is transmitted between generations only by one sex means that if one reconstructs a genealogy of ancestors, only a single line of inheritance needs to be followed. Another type of DNA that shows a uniparental mode of inheritance is mitochondrial DNA (mtDNA), which is transmitted only by females. Thus, the Y-chromosome and mtDNA allow respectively the reconstruction of the male and female lines of inheritance or lineages (Jobling and Tyler-Smith 1995).
2. Variability Of The Y-Chromosome
Although the Y-chromosome determines the male sex, very few genes have been identified on the Y and none of the classical genetic markers are encoded by this chromosome. There is considerable cytogenetic variability of the Y but such variation is difficult to interpret for evolutionary studies. Similarly, most of the variants identified by molecular methods before DHPLC consisted of complex molecular rearrangements that were difficult to characterize and interpret in a population setting. The increasing number of variants that are now being identified can be categorized into two basic types: biallelic and multiallelic. The biallelic markers consist of nucleotide substitution or small insertion deletions while the multiallelic markers generally consist of regions of the DNA in which a variable number of Short Tandem Repeats (STR) of a DNA motif occur (for example a variable number of the tetranucleotide CATG). One basic difference between these two types of markers is that biallelic polymorphisms generally represent single evolutionary events while mutations at the STR loci occur at a relatively higher rate and the mutational process is frequently reversible. This means that identical STR alleles might have a different evolutionary history, as the same allele might be generated by independent mutational events, thus potentially erasing an evolutionary signal when probing too far back in time. However, the combination of these two types of markers actually increases the power of evolutionary analyses based on the Y-chromosome. Slowly evolving biallelic markers can be used to define the basic relationships between Y-chromosomes in an unambiguous form. The rapidly evolving STR markers can then be used to assess the relationships of Y-chromosomes within the basic lineages defined by the biallelic markers. Both kinds of markers can also be used to estimate the age of Y-chromosome lineages based on different evolutionary models, with the consistency of the two estimates thus providing greater confidence in such estimates (Pritchard et al. 1999, Thomson et al. 2000).
3. The Origin And Diversification Of Human Populations
3.1 Human Origins
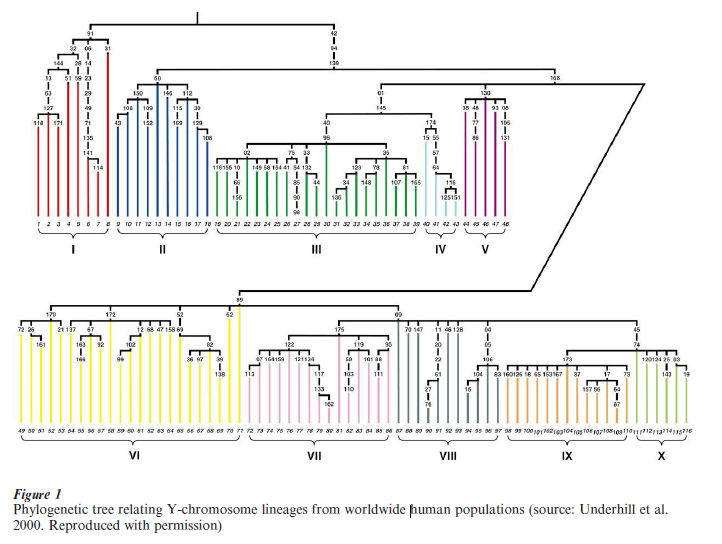
As biallelic markers on the Y-chromosome generally represent events that occurred only once in human evolution it is possible to examine the relatedness of Y-chromosomes worldwide and represent their ancestral relationships on a phylogenetic tree. The analysis of 167 genetic markers in 1,062 males from 21 populations around the world recently allowed the identification of 116 Y-chromosome types (Underhill et al. 2000). Based on the common ancestry of certain types, it is possible to group human Y-chromosomes into ten basic lineages (Fig. 1). Interestingly, the most ancient splits on the tree separate lineages I and II, which are almost exclusively seen in African populations, from all other lineages which have a wider geographic distribution. This observation is in agreement with an origin of human populations in Africa followed by a migration out of that continent of a sample of African men. Descendants of these initial migrants would have founded small populations outside of Africa that would have remained small in size until the end of the last glaciation (around 16,000 years ago) when they would have expanded in numbers, spreading geographically to all inhabitable areas. Such an expansion is clearly evidenced in the Y-chromosome by the large proportion of mutations that are specific to a particular chromosome type. Estimates of the age of Y-chromosome lineages indicate that their initial differentiation is about 59,000 years old with a migration out of Africa around 44,000 years ago (Pritchard et al. 1999, Thomson et al. 2000). These findings are more in agreement with a model of recent African origins of modern humans; however the dates obtained with these data are considerably younger than those estimated from mtDNA and autosomal markers (Horai et al. 1995, Goldstein et al. 1995). This inconsistency might be due to natural selection acting with variable intensity in different genetic systems or could relate to demographic differences between men and women. Overall, the Y-chromosome shows a reduced level of variation when compared with other genetic systems and such variation is highly structured by geography (Seielstad et al. 1998). This makes the Y particularly suitable for the study of the history of specific geographic regions.
3.2 Human Population Diversification
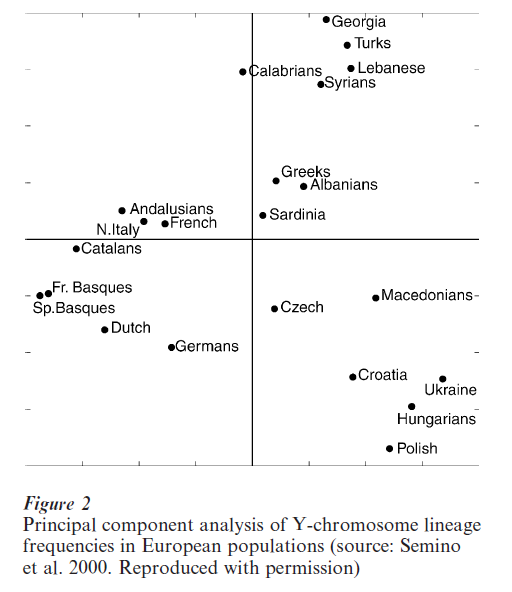
The analysis of Y-chromosome diversity at 22 biallelic markers in 1,007 males from 25 populations of Europe and the Middle East identified 20 different lineages (Semino et al. 2000). However, over 95 percent of the men examined have one of 10 lineages, several of which are closely related. Examining the distribution of these lineages in European populations it was observed that two closely related lineages showed East–West gradients in frequency, with one of them being particularly frequent in the Basque and the other in Eastern Europe (Poland, Hungary, and Ukraine). Phylogenetic and genetic dating analyses suggest that these two types of Y-chromosomes were brought to Europe by people of the Aurignacian culture, which spread to Europe about 35,000 to 40,000 years ago. The East-West gradient is possibly explained by the demographic contraction caused by the last glacial maximum (20,000 to 13,000 years ago) that would have led human populations to seek temporary refuge in more hospitable areas from which they would have subsequently expanded. A third Y-chromosome lineage is about 20,000 years old and is restricted to Europe (where it is seen at highest frequency amongst central Eastern Europeans) although it is closely related to Y-chromosomes frequent in the Middle East. The age and distribution of this lineage suggest that it was introduced by people of the Gravettian culture that appeared in Europe around 20,000 years ago. These people found refuge in central Europe during the last glacial maximum, after which they expanded into Western Europe. Finally, a group of younger and closely related haplotypes show a gradient in frequency from the Middle East into Europe and might have been spread by the people that introduced the practice of agriculture into Europe about 10,000 years ago. Y-chromosome data thus indicate that two Paleolithic and one Neolithic migrations are at the origin of most of the current European population as manifested by three main population clusters seen when lineage frequencies are compared between populations (Fig. 2). The three clusters observed (Western Europe, Eastern Europe, and the Middle East) correspond to the two major glacial refuges and to the area of origin of agriculture.
4. Male And Female Migration In Human Evolution
A very interesting use of genetic data is in teasing apart male and female demographic history throughout human evolution, by comparing population diversity of Y-chromosome, autosomal, and mtDNA markers in native populations from around the world. In an initial evaluation it was observed that the level of genetic variability across populations based on Y-chromosome markers was much greater than that observed with mtDNA markers (Seielstad et al. 1998). This means that the distribution of Y-chromosome variants is geographically more localized than that of mtDNA variants. A possible explanation for this global pattern could be that about 70 percent of human populations practice patrilocality, that is women tend to move into the areas inhabited by the males they marry and consequently move more, on an evolutionary scale, than men. A subsequent refined analysis restricted to South American populations failed to confirm the generality of a higher migration rate of women in native populations (Mesa et al. 2000). A detailed evaluation of the proposal that females have generally had a higher migration rate than males awaits the analysis of more genetic markers and populations. However, it is possible that important regional differences in male / female migration rates might be detected and that these differences relate to the frequency with which patrilocality is practiced in different parts of the world.
5. Maternal And Paternal Origins Of Recently Founded Human Populations
The comparison of mtDNA and Y-chromosome markers is also informative when applied to populations established more recently and such data can be used to refine information obtained from traditional historical sources. Particularly, analyzing different genetic systems can help establish the relative proportion of males and females from particular geographic areas that founded a new population. In the province of Antioquia in Colombia it was seen that about 90 percent of Y-chromosomes were European while 90 percent of the mtDNA lineages were Native American (Carvajal-Carmona et al. 2000). This indicates that the population was founded mostly by European males and Native females. This finding is consistent with the geographic isolation of the province and with historical records indicating that in early colonial times, on average only about 20 percent of the Spanish migrants to the New World were women. However, this result is inconsistent with autosomal markers, which indicate that this population is predominantly of European ancestry. This inconsistency between bi and uniparental markers might relate to the collapse of the Native American population during the first century of Spanish colonization and to the fact that Spanish immigration was a long-term process. These demographic factors might have resulted in a progressive dilution of the founding Native American component as estimated with autosomal markers.
Curiously, the analysis of rapidly evolving STRs indicates that the population of Antioquia seems to have a relatively important Jewish ancestry as evidenced by the finding in this population of Y-chromosome STR lineages seen at high frequency only in Jewish populations. However, no record is available of a Jewish migration to Antioquia. Furthermore, almost simultaneously with the Spanish expansion to America, Jews were expelled from Spain or forced to convert to Christianity. The Y-chromosome findings suggest that amongst the Spanish founders of Antioquia a relative large proportion of converts was represented. No record of such contribution is available probably because the migration to the colonies of converts was considered illegal under colonial rule.
6. Conclusion
Developments resulting from the Human Genome Project have recently catapulted the use of Y-chromosome markers into the forefront of the study of human population origins and diversification. The large number of markers currently available together with novel highly efficient technologies enable analyses of an unprecedented resolution and scale. Evolutionary analyses are facilitated by the fact that slowly evolving markers allow the unambiguous assessment of the evolutionary relationship between Y-chromosomes. Rapidly evolving markers can refine analyses within specific lineages or populations. These studies illuminate not only questions related to the origin of our species and its early diversification but also allow the probing of more recent demographic events. The synthesis of genetic data with information obtained from sources including geology, paleoanthropology, archaeology, and historical demography is allowing a refined reconstruction of human evolution stretching from our origins as a species all the way to the exploration of quite recent historical events.
References:
- Carvajal-Carmona L G, Soto I D, Pineda N, Ortiz-Barrientos D, Duque C, Ospina-Duque J, McCarthy M, Montoya P, Alvarez V M, Bedoya G, Ruiz-Linares A 2000 Strong Amerind White sex bias and a possible sephardic contribution among the founders of a population in Northwest Colombia. American Journal of Human Genetics 67: 1287–95
- Cavalli-Sforza L L, Menozzi P, Piazza A 1994 The History and Geography of Human Genes. Princeton University Press, Princeton, NJ
- Goldstein D B, Ruiz Linares A, Cavalli-Sforza L L, Feldman M W 1995 Genetic absolute dating based on microsatellites and the origin of modern humans. Proceedings of the National Academy of Sciences of the United States of America 92: 6723–7
- Horai S, Hayasaka K, Kondo R, Tsugane K, Takahata N 1995 Recent African origin of modern humans revealed by complete sequences of hominoid mitochondrial DNAs. Proceedings of the National Academy of Sciences of the United States of America 92: 532–6
- Jobling M A, Tyler-Smith C 1995 Fathers and sons: The Y-chromosome and human Trends in Genetics 11: 449–56
- Mesa N R, Mondragon M C, Soto I D, Parra M V, Duque C, Ortiz-Barrientos D, Garcia L F, Velez I D, Bravo M L, Munera J G, Bedoya G, Bortolini M C, Ruiz-Linares A 2000 Autosomal, mtDNA, and Y-Chromosome diversity in Amerinds: Preand post-Columbian patterns of gene flow in South American Journal of Human Genetics 67: 1277–86
- Pritchard J K, Seielstad M T, Perez-Lezaun A, Feldman M W 1999 Population growth of human Y-chromosomes: A study of Y-chromosome Molecular Biology and Evolution 16: 1791–8
- Ruiz-Linares A 1999 Microsatellites and the reconstruction of the history of human In: Goldstein D B, Schlotterer C (eds.) Microsatellites. Evolution and Applications. Oxford University Press, Oxford, pp. 183–97
- Semino O, Passarino G, Oefner P J, Lin A A, Arbuzova S, Beckman L E, De Benedictis G, Francalacci P, Kouvatsi A, Limborska S, Marcikiae M, Mika A, Mika B, Primorac D, Santachiara-Benerecetti A S, Cavalli-Sforza L L, Underhill P A 2000 The genetic legacy of Palaeolithic homo sapiens sapiens in extant Europeans: A Y-chromosome perspective. Science 290: 1155–9
- Seielstad M T, Minch E, Cavalli-Sforza L L 1998 Genetic Evidence for a higher female migration rate in humans. Nature Genetics 20: 278–80
- Thomson R, Pritchard J K, Shen P, Oefner P J, Feldman M W 2000 Recent common ancestry of human Y-chromosomes: Evidence from DNA sequence Proceedings of the National Academy of Sciences 97: 7360–5
- Underhill P A, Shen P, Lin A A, Jin L, Passarino G, Yang W H, Kauffman E, Bonne-Tamir B, Bertranpetit J, Francalacci P, Ibrahim M, Jenkins T, Kidd J R, Mehdi S Q, Seielstad M T, Wells R S, Piazza A, Davis R W, Feldman M W, CavalliSforza L L, Oefner P J 2000 Y-chromosome sequence variation and the history of human Nature Genetics 26: 358–61.




